 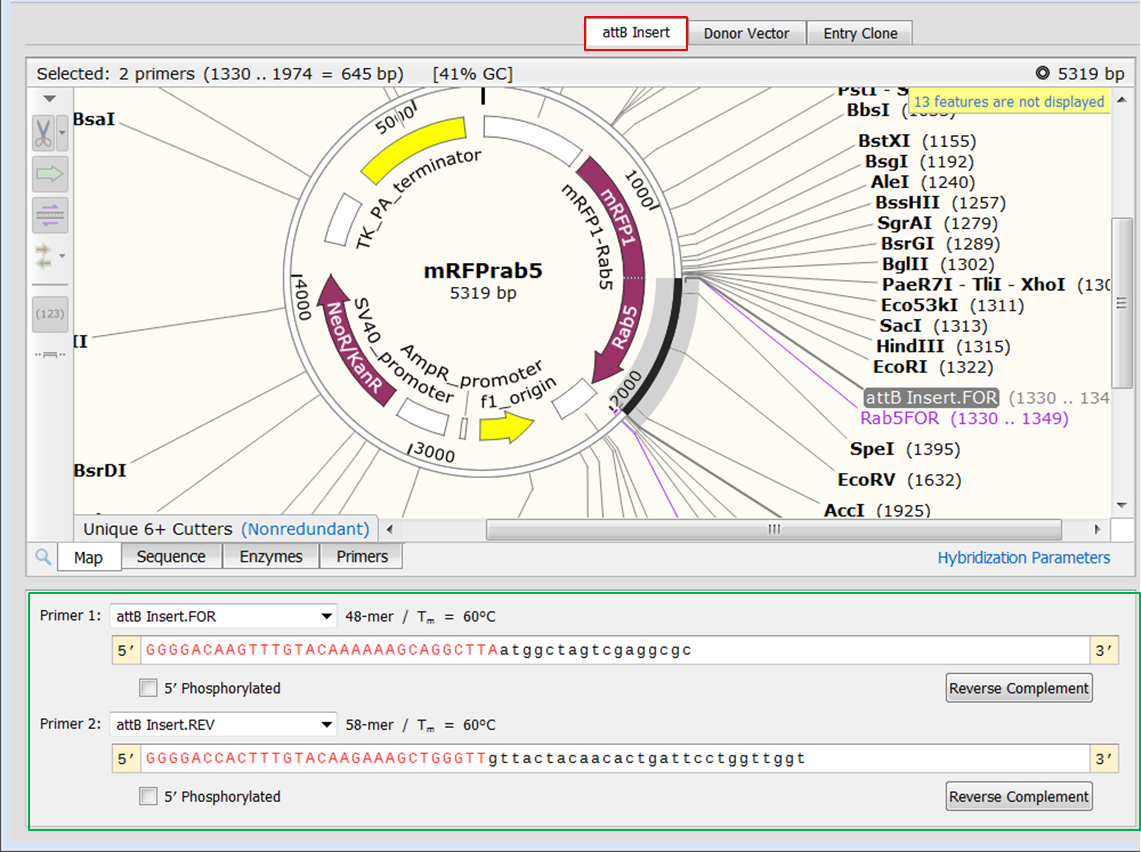
Try and make the process as simple as possible, so that the actual cloning is easy as well. If at all possible, use enzymes that only cut once within a vector, leave sticky overhangs (that is 5′ or 3′ DNA overhangs), and have simple recognition sites (usually 6-8 base pair sites). It is important to consider how many times the enzyme cuts in the vector, the overhang that is left behind, and the type of enzyme recognition site. For many cloning experiments, this can be complicated. I was only concerned with removing the old guide RNA sequences and replacing them with those that I had designed:įor restriction cloning to work, you need to find enzymes that cut both the target vector and the incoming DNA, so you need to examine the cut sites. The vector consists of the old guides, a plant ubiquitin promoter to express the Cas9, a selection gene, and various plasmid maintenance genes. The first file to look at is the destination vector, so I opened mine up: I use SnapGene, but there are a large variety of programs out there that have these capabilities. The first thing to do is open up your sequence files in your program of choice. Pull Up Your Target Vector, Desired Insert, and Other DNA Sequences on the Computer Screen I also knew the new guides were being synthesized and shipped to me in a standard vector, pUC19. I knew that I didn’t have to worry about anything being in frame, because that was already done for me. This seems simple, but it is the most important part of the planning process.įor my cloning experiment I needed to remove the guide RNA sequences in a Cas9 vector that I had, and replace them with a new set of sequences I had synthesized by a company. Make sure to make note of things like frame requirements, etc. Write down exactly which vector you have, what components you need where, and any other stipulations that are required at the end of the process. Lay Out What You Have and What You Need for Restriction Cloning It had a little of everything, so it will cover most of the problems you will run into in your cloning endeavors. We are here to help! The easiest way to learn anything is to do it, so I thought I would take you step-by-step through the last cloning set-up I did. If you are just starting with restriction cloning, it can be a bit of a struggle to learn the process, but that is ok. 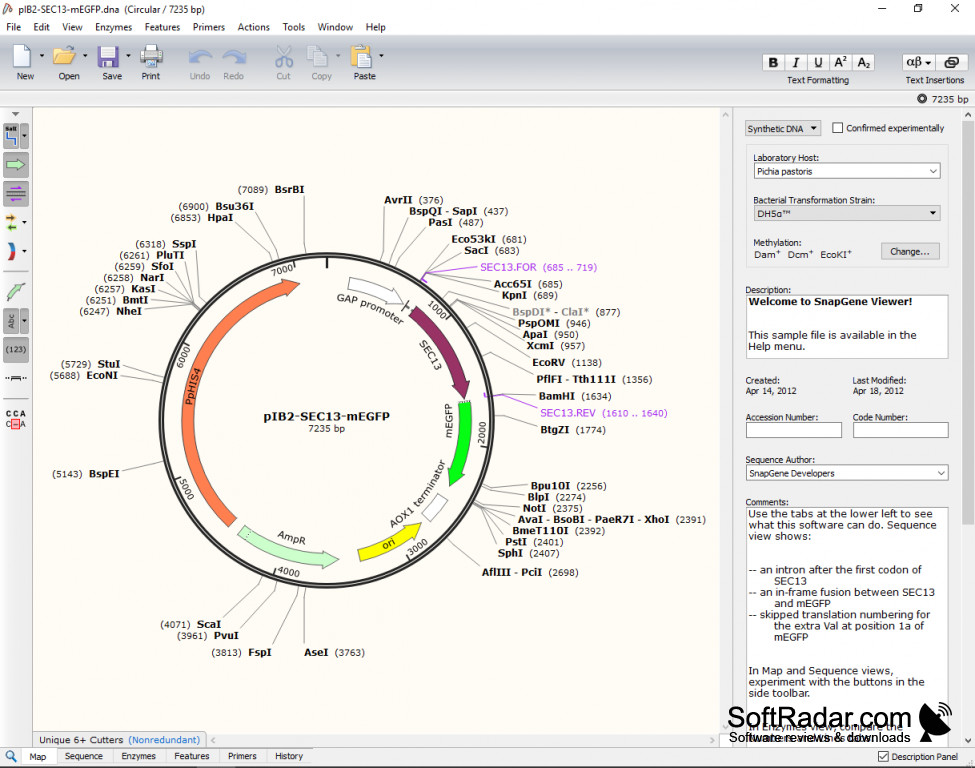
While getting each of the steps correct can be a bit of a hassle, in my mind, the most difficult part of the whole process is planning it. You simply cut the target vector and insert with the same enzymes, clean digested vector and insert up, ligate the two together, transform the ligated vector and insert into bacteria, and then screen. Gene editing for cancer therapy/drug discoveryĭSS Takara Bio India Pvt.Restriction cloning, at its core, is quite simple. Immunizing mice and optimizing vaccine targets Webinar: Speeding up diagnostic developmentĬharacterizing the viral genome and host response Guest webinar: developing and validating molecular diagnostic tests Guest webinar: extraction-free SARS-CoV-2 detection Phosphoprotein and glycoprotein purification SmartChip Real-Time PCR System introduction Outsourcing stem cell-based disease model development Gene and cell therapy manufacturing servicesĮxploring OEM and custom enzyme partnerships RNA extraction and analysis for real-time qPCR Primary antibodies and ELISAs by research area IDimerize inducible protein interaction systemsĬustom business friendly and automation-ready solutions NOT FOR USE IN DIAGNOSTIC PROCEDURES (EXCEPT AS SPECIFICALLY NOTED). Our mission is to develop high-quality innovative tools and services to accelerate discovery.įOR RESEARCH USE ONLY. As a member of the Takara Bio Group, TBUSA is part of a company that holds a leadership position in the global market and is committed to improving the human condition through biotechnology. provides kits, reagents, instruments, and services that help researchers explore questions about gene discovery, regulation, and function. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
